Learning Objective
By the end of this section, you will be able to:
- Describe the structure of DNA
- Describe how eukaryotic and prokaryotic DNA is arranged in the cell
In the 1950s, Francis Crick and James Watson worked together at the University of Cambridge, England, to determine the structure of DNA. Other scientists, such as Linus Pauling and Maurice Wilkins, were also actively exploring this field. Pauling had discovered the secondary structure of proteins using X-ray crystallography. X-ray crystallography is a method for investigating molecular structure by observing the patterns formed by X-rays shot through a crystal of the substance. The patterns give important information about the structure of the molecule of interest. In Wilkins’ lab, researcher Rosalind Franklin was using X-ray crystallography to understand the structure of DNA. Watson and Crick were able to piece together the puzzle of the DNA molecule using Franklin's data (Figure 9.2). Watson and Crick also had key pieces of information available from other researchers such as Chargaff’s rules. Chargaff had shown that of the four kinds of monomers (nucleotides) present in a DNA molecule, two types were always present in equal amounts and the remaining two types were also always present in equal amounts. This meant they were always paired in some way. In 1962, James Watson, Francis Crick, and Maurice Wilkins were awarded the Nobel Prize in Medicine for their work in determining the structure of DNA.

Now let’s consider the structure of the two types of nucleic acids, deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). The building blocks of DNA are nucleotides, which are made up of three parts: a deoxyribose (5-carbon sugar), a phosphate group, and a nitrogenous base (Figure 9.3). There are four types of nitrogenous bases in DNA. Adenine (A) and guanine (G) are double-ringed purines, and cytosine (C) and thymine (T) are smaller, single-ringed pyrimidines. The nucleotide is named according to the nitrogenous base it contains.
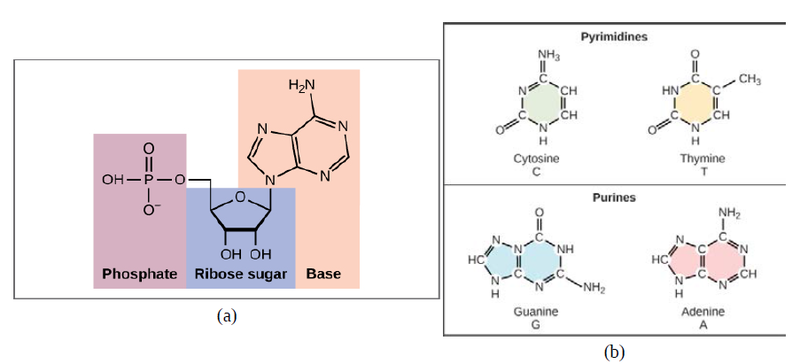
The phosphate group of one nucleotide bonds covalently with the sugar molecule of the next nucleotide, and so on, forming a long polymer of nucleotide monomers. The sugar–phosphate groups line up in a “backbone” for each single strand of DNA, and the nucleotide bases stick out from this backbone. The carbon atoms of the five-carbon sugar are numbered clockwise from the oxygen as 1', 2', 3', 4', and 5' (1' is read as “one prime”). The phosphate group is attached to the 5' carbon of one nucleotide and the 3' carbon of the next nucleotide. In its natural state, each DNA molecule is actually composed of two single strands held together along their length with hydrogen bonds between the bases.
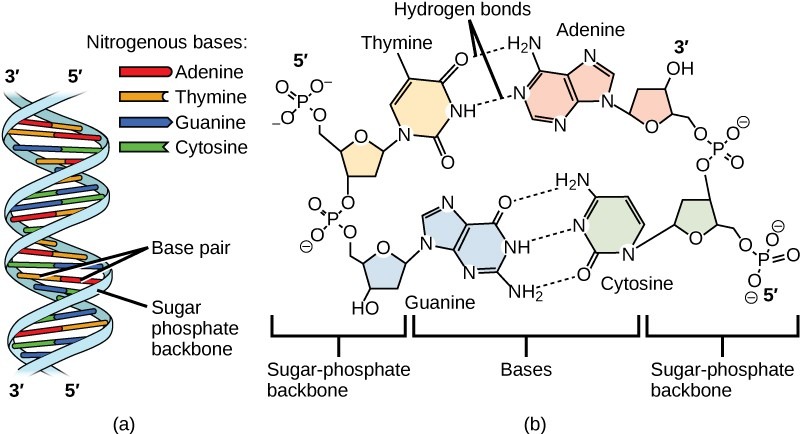
Watson and Crick proposed that the DNA is made up of two strands that are twisted around each other to form a right-handed helix, called a double helix. Base-pairing takes place between a purine and pyrimidine: namely, A pairs with T, and G pairs with C. In other words, adenine and thymine are complementary base pairs, and cytosine and guanine are also complementary base pairs. This is the basis for Chargaff’s rule; because of their complementarity, there is as much adenine as thymine in a DNA molecule and as much guanine as cytosine. Adenine and thymine are connected by two hydrogen bonds, and cytosine and guanine are connected by three hydrogen bonds. The two strands are anti-parallel in nature; that is, one strand will have the 3' carbon of the sugar in the “upward” position, whereas the other strand will have the 5' carbon in the upward position. The diameter of the DNA double helix is uniform throughout because a purine (two rings) always pairs with a pyrimidine (one ring) and their combined lengths are always equal. (Figure 9.4).
- 6364 reads






