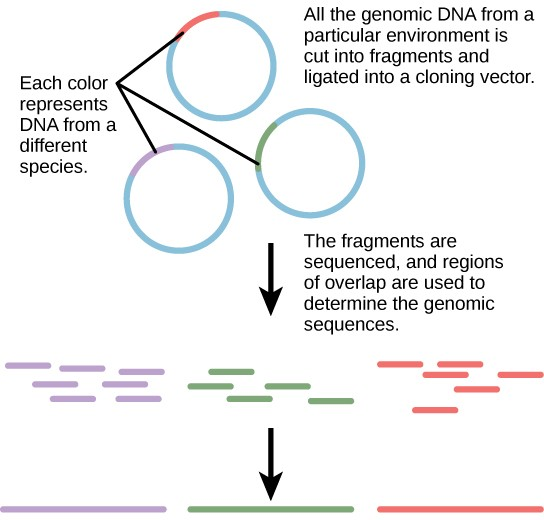
Traditionally, microbiology has been taught with the view that microorganisms are best studied under pure culture conditions, which involves isolating a single type of cell and culturing it in the laboratory. Because microorganisms can go through several generations in a matter of hours, their gene expression profiles adapt to the new laboratory environment very quickly. On the other hand, many species resist being cultured in isolation. Most microorganisms do not live as isolated entities, but in microbial communities known as biofilms. For all of these reasons, pure culture is not always the best way to study microorganisms. Metagenomics is the study of the collective genomes of multiple species that grow and interact in an environmental niche. Metagenomics can be used to identify new species more rapidly and to analyze the effect of pollutants on the environment (Figure 10.13). Metagenomics techniques can now also be applied to communities of higher eukaryotes, such as fish.
- 2199 reads






